 $$\begin\) being the average of voxel values from the enhanced image and \(stdev\) being the standard deviation from the same image (Fig. A thresholding is then applied on the image to segment the final objects (Fig. The voxel values of the enhanced image are used to compute a threshold value ( \(t\)). NODeJ assumes one nucleus per image and uses as an input the raw image of the nucleus as well as the mask of this nucleus (binary image of the nucleus) and computes the enhanced image resulting from the Laplacian operator ( \(\delta v_x\)). All of these methods use the distribution of voxel values to define the connected components of an image. Other methods also included in this group are the watershed and the Isodata algorithms. The Laplacian method belongs to a group of mathematical methods for automated segmentation of objects based on the distribution of voxel intensities across an image. Our method is based on a Laplacian algorithm for object boundary detection. The program can be run as an ImageJ plugin through the graphic user interface (GUI) or the command line (CLI) mode to handle large datasets. NODeJ can be used to process images of nuclei from samples expressing fluorescent reporters, or from fixed tissues or isolated nuclei stained with DNA dyes. The relevant parameters for the objects detected in the raw images are computed using NucleusJ2.0 methods implemented within NODeJ. 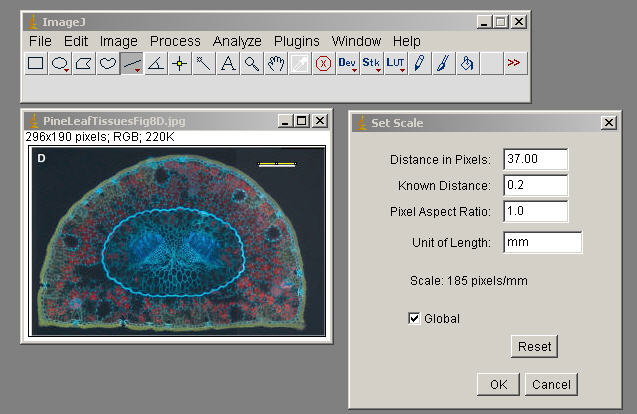
This results in an increase in the contrast of objects of interest and allows to define a threshold on the enhanced image to obtain the segmented objects. 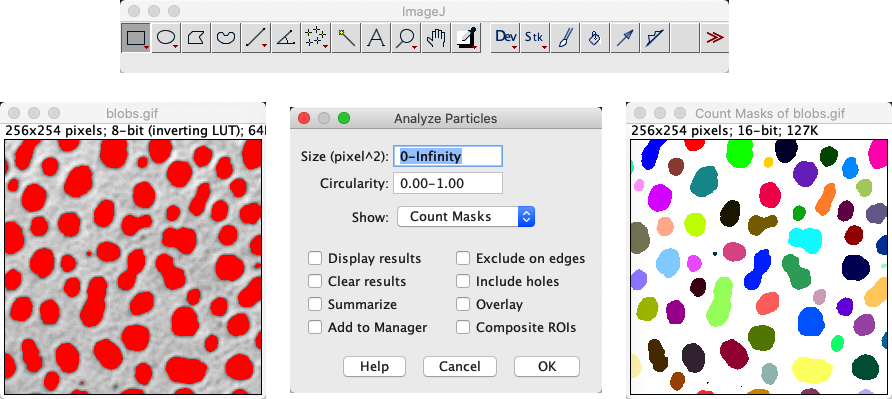
NODeJ implements an algorithm based on a Laplacian convolution. Here, we describe Nuclear Object DetectionJ (NODeJ), a new tool to automatically segment subnuclear objects such as chromocenters and Fluorescence In Situ Hybridization (FISH) signals. This induces limitations for high-throughput data analysis and for achieving consistency when processing a large number of images, while also potentially introducing user bias. While NucleusJ2.0 can automatically detect nuclei in 3D images and compute their general characteristics, further segmentation of subnuclear structures is at best a semi-automated procedure that requires user input. shape and size), as well as chromatin organization. We previously developed a workflow called NucleusJ2.0, designed to compute nuclear morphometric parameters (e.g. Three-dimensional (3D) imaging methods are widely used to investigate nuclear morphology. Changes in the location of heterochromatic domains (also known as chromocenters in Arabidopsis thaliana ) can have a drastic effect on gene expression and on the maintenance of heterochromatin itself. The spatial arrangement of chromatin within the nucleus has fundamental consequences for the accessibility and activity of regions of the genome. The nucleus is a dynamic and complex structure that changes morphology and organization of its DNA content during development. The images used in this study are publicly available at and. NODeJ can be downloaded from with source code, documentation and further information avaliable at. NODeJ is written in Java and provided as an ImageJ plugin with a command line option to perform more high-throughput analyses. NODeJ allows for efficient automated analysis of subnuclear structures by avoiding the semi-automated steps, resulting in reduced processing time and analytical bias. NODeJ is also able to detect signals in nuclei from DNA FISH experiments, allowing for the analysis of specific targets of interest. We reanalyzed public datasets and determined that NODeJ is able to accurately identify heterochromatin domains from a diverse set of Arabidopsis thaliana nuclei stained with DAPI or Hoechst. NODeJ performs a Laplacian convolution on the mask of a nucleus to enhance the contrast of intra-nuclear objects and allow their detection. This program segments and analyzes high intensity domains in nuclei from 3D images. 
We present an automated method, Nuclear Object DetectionJ (NODeJ), developed as an imageJ plugin. Three-dimensional imaging technology allows to follow these dynamic changes, but only a few semi-automated processing methods currently exist for quantitative analysis of the 3D chromatin organization. Chromatin organization is a dynamic process that can be affected by biotic and abiotic stresses. The three-dimensional nuclear arrangement of chromatin impacts many cellular processes operating at the DNA level in animal and plant systems. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
